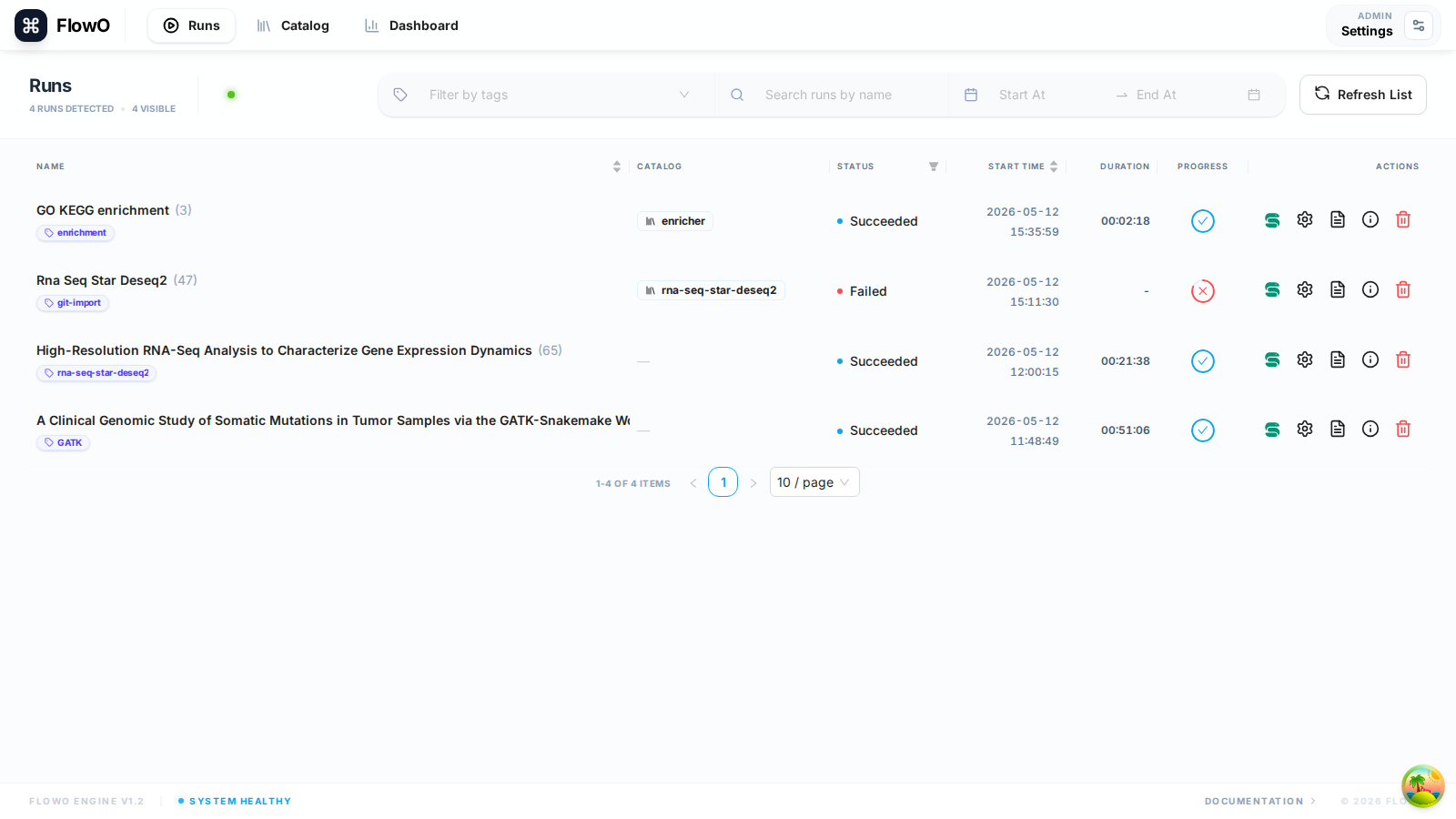
Run Snakemake with FlowO
Integrating FlowO into your existing Snakemake execution is as simple as adding a single flag.
The --logger Flag
To enable FlowO reporting, append the --logger flowo argument to your Snakemake command:
snakemake --logger flowo
Customizing the Run
You can provide additional metadata to help organize your runs in the dashboard using logger-specific arguments.
Set a Custom Name
By default, FlowO uses the project directory name. Use --logger-flowo-name to set a more descriptive title.
snakemake --logger flowo --logger-flowo-name "RNA-Seq Sample Analysis"
Add Tags
Use comma-separated tags with --logger-flowo-tags to categorize your runs (e.g., project IDs, sample types, or experimental conditions).
snakemake --logger flowo --logger-flowo-tags "project-x,rna-seq,control"
Associate with a Catalog Entry
If you are running a workflow derived from the FlowO Catalog, specify the slug using --logger-flowo-catalog.
snakemake --logger flowo --logger-flowo-catalog "rna-seq-pipeline"
Configuration via Snakefile/Config
Instead of long command-line arguments, you can define FlowO settings directly in your Snakemake configuration.
In config.yaml:
flowo_project_name: "My Custom Project"
flowo_tags: "experiment-1,debug"
In a Python dict inside the Snakefile:
config["flowo_project_name"] = "My Custom Project"
config["flowo_tags"] = "experiment-1,debug"
Verification
When the logger is active, switch to Runs (/runs) in the browser: the new row should appear within a second or two while jobs stream in over SSE.